The evolution of RNA aptamers has taken a significant leap forward with the introduction of GRAPE-LM, a groundbreaking generative artificial intelligence framework. This new system promises to streamline the traditionally labor-intensive process of aptamer discovery, allowing for the effective evolution of RNA in a single round rather than the previously required multi-round screenings.
Historically, the directed evolution of biomolecules has been an iterative and time-consuming process. While advancements in artificial intelligence have accelerated protein evolution, RNA remains challenging to evolve efficiently. RNA aptamers, known for their specific binding properties, present an ideal target for this innovation. Traditional methods often depend on extensive, multi-phase screening, a factor that GRAPE-LM aims to overcome.
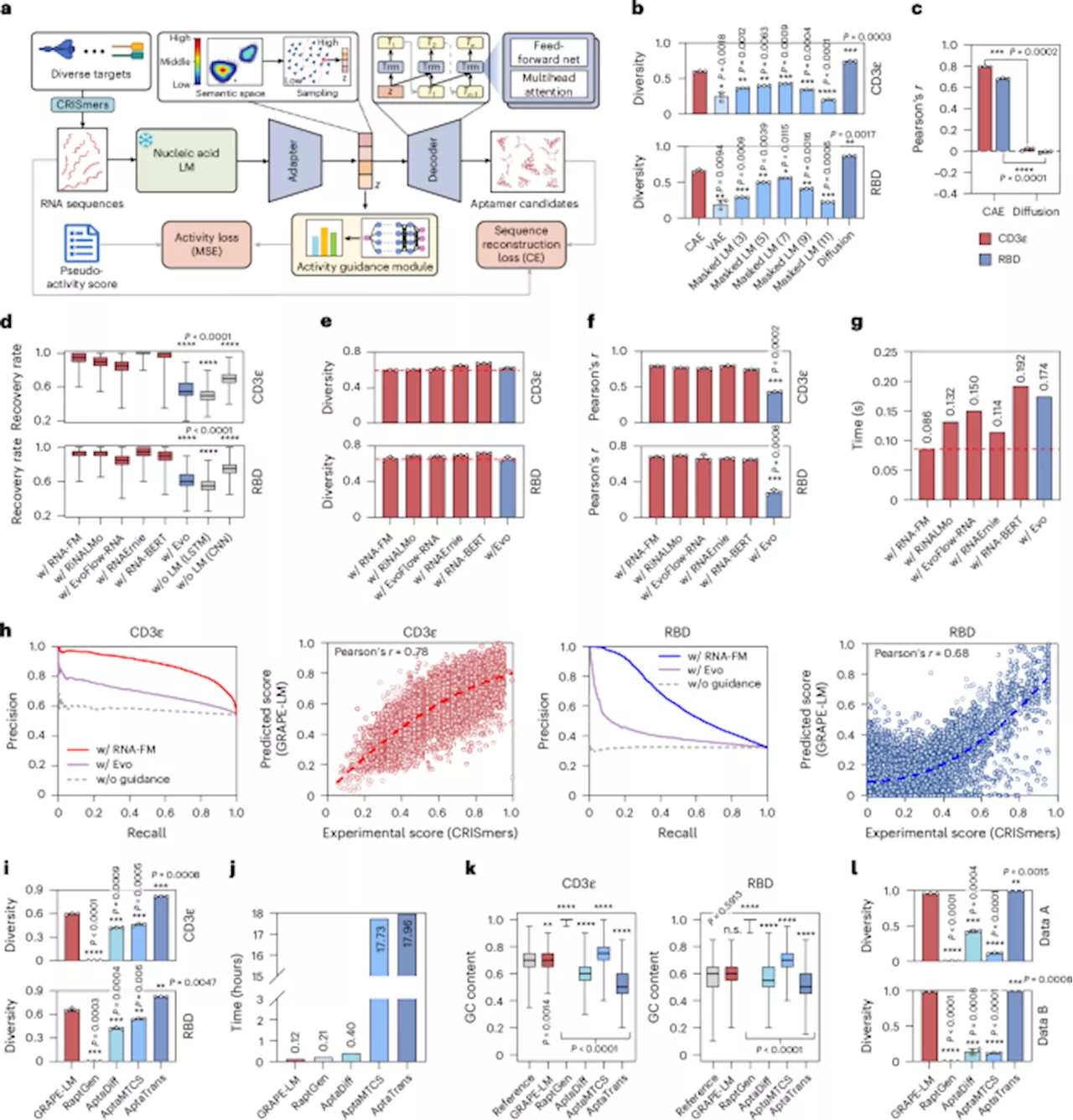
GRAPE-LM employs a transformer-based conditional autoencoder combined with nucleic acid language models. It utilizes data from CRISPR-Cas-based aptamer screening derived from intracellular environments. This novel approach enables the framework to identify high-performance RNA aptamers based on just one round of screening data, significantly reducing the time and resources required for development.
To validate its effectiveness, researchers tested GRAPE-LM on three distinct targets: the human T cell receptor CD3ε, the receptor-binding domain of the SARS-CoV-2 spike protein, and the human oncogenic transcription factor c-Myc. Remarkably, the RNA aptamers generated by GRAPE-LM outperformed those developed through multiple rounds of human selection and optimization.
The results from these validation tests underscore the potential of GRAPE-LM to transform the landscape of RNA aptamer development. By leveraging a single round of CRISPR-Cas-based screening, the framework not only accelerates the process but also enhances the quality of the aptamers produced.
This innovation holds particular promise for the fields of diagnostics and therapeutics, where RNA aptamers can be employed to selectively bind and neutralize specific targets, including viral proteins and cancer markers. The ability to quickly generate high-affinity aptamers could significantly impact the speed at which new treatments and diagnostic tools are developed.
The research was conducted by a team from the Shenzhen Institute of Advanced Technology and supported by various funding sources, including the National Natural Science Foundation of China and local government programs. The collaborative effort highlights the importance of interdisciplinary approaches in tackling complex scientific challenges.
In summary, the introduction of GRAPE-LM marks a pivotal advancement in the evolution of RNA aptamers. By enabling the efficient generation of functional aptamers in a single round, this technology not only saves time but also enhances the potential for innovative applications in medicine and biotechnology. As research continues, it will be essential to monitor how this framework can be adapted and applied across various biological systems and therapeutic targets.